Related Resources
Insights
Dichroic Mirrors and Filters for SPECTRA, CELESTA, and ZIVA Light Engines
Insights
Precise Power Regulation on each of Lumencor’s Newest and Brightest Light Engines: AURA, SPECTRA, CELESTA, and ZIVA Light Engines
Insights
Recommended Operating Conditions for CELESTA, SPECTRA and ZIVA Light Engines
Insights



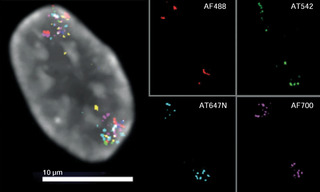
![Figure 1. Localization of 16 FISH probes in a single retinal pigment epithelial (RPE) cell nucleus. Individual images for each of the 6 fluorophores used to construct the combinatorially color encoded composite image are shown in the panels on the right. Gray represents DNA stained with Hoechst 33342. Reproduced from [2] under CC BY 4.0.](https://cms.lumencor.com/system/uploads/fae/image/asset/955/xs_Figure_1.jpg)
![Figure 2. Spectral output of customized SPECTRA light engine for single-molecule FISH localization of 16 DNA loci [2].](https://cms.lumencor.com/system/uploads/fae/image/asset/956/xs_Figure_2.jpg)