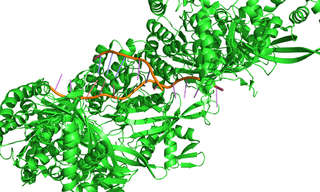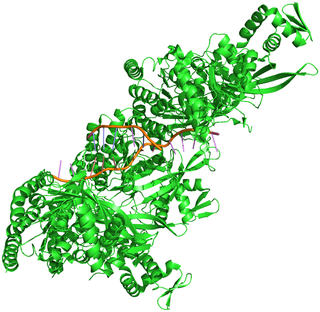
Figure 1. Crystal structure of the RecA-DNA complex
Efficient repair of double-strand breaks (DSBs) in genomic DNA is critical in protecting cells from potentially fatal chromosomal aberrations. Fast and accurate repair of DSBs is achieved by homologous recombination, in which the recombinase enzyme RecA forms a nucleoprotein filament with the single-stranded DNA present at a DNA break and searches for an undamaged homologous dsDNA segment to use as a template for break repair (Figure 1). To understand how the spatiotemporal dynamics of homologous recombination facilitate the speed and accuracy of the process, researchers from the Science for Life Laboratory at Uppsala University relied on solid-state illumination from a SPECTRA Light Engine to visualize the repair of DSBs in single Escherichia coli cells by high-throughput multicolor fluorescence microscopy [1]. Cells undergoing DSB repair were identified on the basis of enhanced CFP expression coupled to the DNA damage-triggered SOS response. RecA activity during DSB repair was visualized using a RecA–YFP fusion, with mCherry–tagged chromosome-partitioning protein ParB marking chromosome break locations. Image analysis revealed that repair of DSBs between segregated sister loci is completed in 15 ± 5 minutes and that dsDNA homology search takes less than 9 ± 3 minutes. This result implies that homology pairing occurs via radial migration of a segment of chromosomal dsDNA to the RecA–ssDNA filament, rather than 3-dimensional diffusion of two segments in the whole cell.