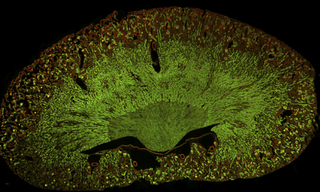
Integration of pooled CRISPR genetic screening with cellular and sub-cellular imaging readouts is critical to improving phenotypic definition in image-based genetic knockout studies. In a recent paper published in Nature Methods[1], Wheeler and co-workers describe screening CRISPR-infected HEK293T cells on microraft arrays, followed by automated high-resolution confocal imaging to identify regulators of stress granules, which are punctate protein–RNA assemblies that form during cellular stress. Microraft arrays (commercially available from Cell Microsystems; Durham, NC) are an attractive platform for screening bulk-infected cells because thousands of clonal cell colonies (~5–20 cells per colony) can be cultured in isolation from one another. Confocal imaging of microraft arrays was accomplished using a CELESTA Light Engine coupled to a CrestOptics X-Light V2 L-FOV spinning disk confocal. The 405 nm, 520 nm, 546 nm, and 638 nm lines of the CELESTA light source were used to excite fluorescence of DAPI, mCitrine, mCherry, and Alexa Fluor 633 respectively. The screen identified and validated six previously known stress granule modulators, along with 17 RNA-binding proteins (RBPs) that, when depleted, reduce sodium arsenite-induced stress granules in human cells.
- Jul 28, 2020